The recovery of silver from ores by hydrometallurgical techniques changed considerably toward the end of the 19th and in the early part of the 20th century. Metallurgical techniques, such as amalgamation, chloride roasting followed by amalgamation, hyposulphite leaching, and the Patio process (amalgamation of silver formed by the reaction of silver minerals with sodium chloride and copper sulfate) have been replaced by cyanidation. By the turn of the century, cyanidation was used almost exclusively, except amalgamation was used in placer operations. Many of the tailings from the old processes have been reworked by the cyanidation technique. However, not all silver ores are readily amenable to cyanidation and those that are not present a serious problem to the mill operator. The extraction of silver by cyanidation can be seriously deterred by manganese minerals in the form of manganoargentojarosites, oxides, or carbonates of manganese by various oxidized and reduced iron minerals, as well as a host of cyanicides. With certain ores, silver extraction can be improved by finer grinding to liberate the silver; however, finer grinding is not universally effective and is frequently limited by the costs involved.
The Bureau of Mines undertook a study of refractory silver ores to determine the mineral assemblage of the ores and the effects that alteration of these minerals might have on extraction of silver by cyanide, chlorine, sulfuric acid, and sulfur dioxide in brine solution. This report presents data obtained from leaching studies on refractory silver ores and dump material. Emphasis was placed on alteration of iron and manganese minerals by electrooxidation and sulfurous acid leaching and on determination of the effect on silver extraction. The electrooxidation technique was developed by the Bureau of Mines for treatment of carbonaceous gold ore, cinnabar ore, and molybdenum-rhenium ore and concentrates. The process involves passing current through a brine-pulp to generate chlorine at the anode and base at the cathode. The chlorine and base combine to form hypochlorite which then oxidizes the carbonaceous and sulfide portions of the ore.
Description of the Ore
The ores used for this study were obtained from the Candelaria Mining District in west central Nevada and the Round Mountain District of Colorado. The silver, iron, and manganese content of the ores used in this investigation is shown in table 1. Mineralogy studies of low-grade ore collected from the Mount Diablo dump in the Candelaria Mining District showed a heterogeneous mixture of several rock types. Included were fine-grained shales and argellites, acidic and intermediate dike rocks, hydrothermally altered material from both of the previous types, and oxidized vein material thought to contain most of the silver values. All the ores were similar in mineral composition, except that the ore from Round Mountain contained appreciably more manganese minerals.
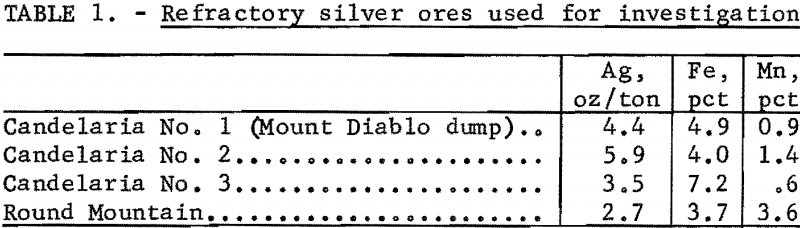
Because of the wide range of rock types present, along with the low overall silver content (2.7 to 5.9 ounces Ag per ton), a sample of high-grade oxidized vein material was obtained from the Mount Diablo dump for initial studies. Changes in silver content resulting from metallurgical treatment of a particular mineral in the high-grade sample were easier to monitor, and specific silver-bearing phases left after treatment could be identified. The high-grade sample (about 40 ounces silver per ton) consisted mainly of quartz, iron and manganese oxides, and jarosite-type minerals. A combination of ore microscopy and electron microprobe examinations revealed that silver occurred in several distinct phases. Most readily visible were large areas of extremely fine-grained mixtures of jarosite group minerals including potassium, lead, and silver-bearing varieties [X’Fe6(OH)12(SO4)4 X’ = Kg , Ag2, Pb]. Silver was concentrated within discrete grains, which were usually surrounded by quartz gangue. The host minerals included oxidized iron-sulfides, antimony-iron oxides, manganese-lead oxides, and antimony-rich sulfides. Well-defined rims of silver sulfide on pyrite were also present. These rims were often several microns wide and usually rimmed by antimony sulfide.
The sample of high-grade material used for the metallurgical studies was prepared in the following manner: The ore was ground and screened. All screen fractions contained approximately the same amount of silver; however, the minus 100 plus 500 screen fraction contained the largest concentration of heavy minerals, iron oxides, and manganese (IV) and (II) oxide-carbonates. To further upgrade the silver minerals in the minus 100 plus 500 fraction, a heavy liquid separation was made. The heavy minerals were concentrated in a fraction with a density greater than 3.30. Composition of this concentrate was 35 percent iron, 0.75 percent manganese, and 50 ounces of silver per ton.
Apparatus and Analytical Procedure
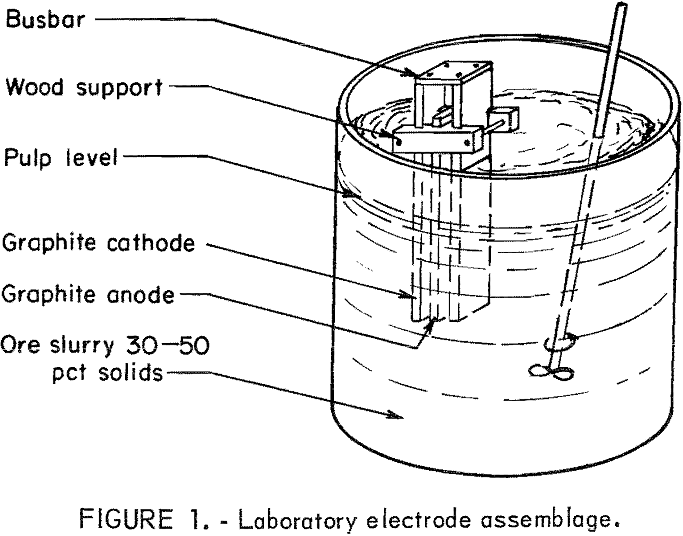
Electrooxidation experiments were conducted using a 20-percent sodium chloride solution and a pulp density of 35 percent. The cell used for experiments consisted of three carbon electrodes suspended in the pulp as shown in figure 1. The current density was held constant at 0.5 amp per in². Current was supplied by a constant amperage dc rectifier.
Sulfurous acid and sulfuric acid experiments were conducted in open beakers in which the ore slurry was stirred constantly, or in bottles placed on rolls (standard bottle test). The source of sulfur dioxide was an aqueous solution of 6 percent H2SO3. No attempt was made to control gas pressure above the liquid for these experiments.
Sulfur dioxide gas experiments were conducted by bubbling a sulfur dioxide-air mixture into a stirred pulp in an open beaker. The SO2 content of the gas mixture was 12 percent by volume, a concentration normally obtained when sulfur is burned to produce sulfur dioxide.
Silver was determined in solid samples by the standard fire assay technique; iron and manganese were determined by
wet-chemical methods. Silver, iron, and manganese in solution samples were determined by atomic absorption.
Cyanidation tests were conducted using 8 pounds of cyanide per ton of solution, a pulp density of 40 percent, ambient room temperature, and a digestion time of 24 hours. Lime was used to maintain a pH of approximately 12 for these tests.
Results and Discussion
Initial experiments were conducted using the minus 100 plus 150 fraction of the high-grade ore concentrate to determine the relationship between alteration of the iron and manganese minerals and silver extraction. The minus 100 plus 150 fraction was used because smaller particle sizes are difficult to mount for electron microprobe examination. Initially, the material was electrooxidized to determine the extent to which silver could be extracted from the iron, manganese, and jarosite minerals by oxidation. The silver which occurred as sulfide mineral was converted to silver chloride in the oxidizing brine solution,
Ag2S + 4OCl- → 2AgCl + SO4= + 2Cl-…………………………………………..(1)
Silver chloride would be soluble in either cyanide solution as cyanide complex or brine solution containing excess chloride ion as the tetrachloro complex,
AgCl + 2CN- → Ag(CN)2- + Cl-……………………………………………(2)
AgCl + 3Cl- → AgCl4-³……………………………………………………(3)
In a series of electrooxidation experiments, the maximum silver extraction was 34 percent, even though 192 kilowatt-hours per ton of ore was used. Only minor amounts of manganese and iron dissolved during the treatment. Cyanidation of the electrolyzed tails did not increase silver extraction. The results indicated that silver minerals, other than sulfides, were not altered sufficiently by electrooxidation to allow the silver to react with chloride or cyanide ions.
Mineralogy studies have shown that the manganese mineral contains a mixture of Mn(IV) and Mn(II). The usual minerals for these valences are MnO2 and MnO, respectively. A reducing acid is required to dissolve Mn(IV), whereas a strong acid such as H2SO4 will dissolve Mn(II). An acidic system which provides both a reducing and a strong acid is a sulfur dioxide-water (sulfurous acid) leach. The reactions for such a system are shown in equations 4 through 6:
MnO2 (s) + SO2 (g) → MnSO4 (aq)…………………………………………….(4)
H2O + SO2(g) + ½O2(g) → H2SO4 (aq)……………………………………….(5)
MnO + H2SO4 → MnSO4 + H2O………………………………….(6)
The oxide iron minerals would also be soluble in an acidic media,
Fe2O3 + 6H+ → 2Fe+³ + 3H2O……………………………………………….(7)
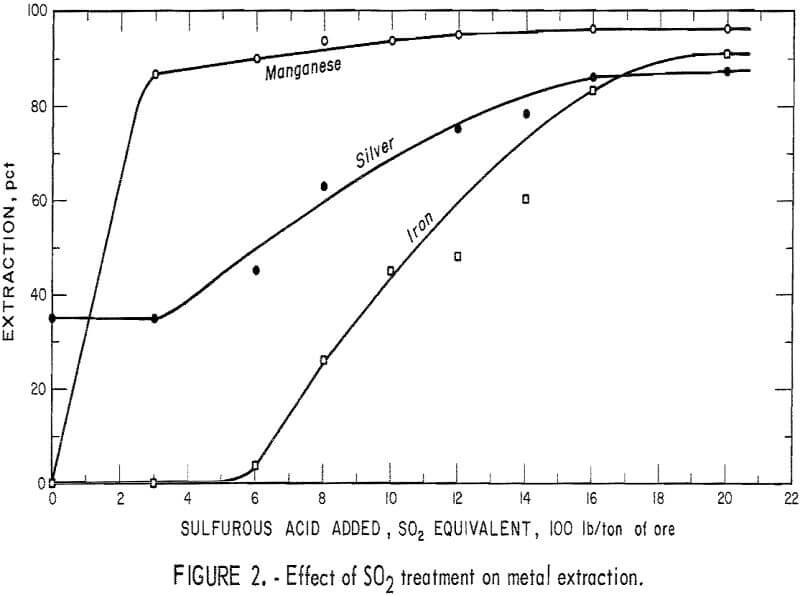
A series of sulfurous acid-leaching experiments was conducted to determine the effect of iron and manganese removal on silver extraction. The ore was leached with H2SO3, and the filtered pulp was treated by cyanidation. The iron and manganese dissolved during the acid treatment, as chloride complex salts, liberating the silver. The data in figure 2 show that the manganese minerals consumed essentially all of the initial 300 pounds of SO2 added to the system and that the iron extraction was nil. Corresponding silver extraction remained constant at the 35-percent level obtained by direct cyanidation of the untreated ore. The iron and silver minerals started to dissolve on further addition of sulfur dioxide. The iron and silver codissolved with increasing addition of sulfur dioxide until silver extraction leveled off at 86 percent. The data indicated that most of the silver was contained in the limonitic iron minerals. Analysis of the tails after treatment with sulfur dioxide and cyanide leaching showed that the silver remaining in the tails was in the form of the argentojarosite [Ag2Fe (OH)12 (S04 )4] mineral.
Additional treatment of the tails with 3 N HCl, H2SO4 , or HNO3 followed by cyanidation of the residues resulted in increasing silver extraction to 95 percent. The high silver extractions indicated that the high concentration of acid destroyed the argentojarosite mineral and that very little silver was encapsulated in a silicious matrix. The treatment with strong acid would have no commercial application, because acid consumption was 800 to 1,000 pounds per ton.
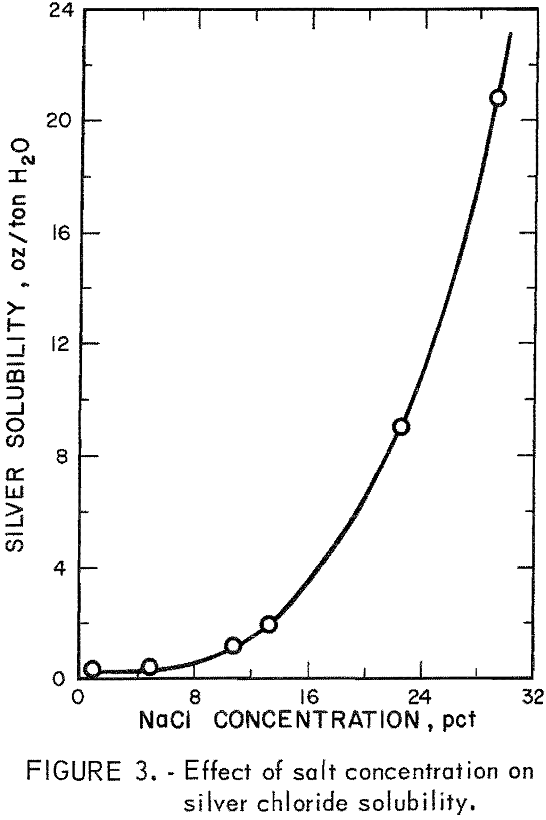
The low-grade refractory silver ore was studied in subsequent experiments using data obtained from baseline experiments on selected high-grade silver ore as guidelines. Eighty-seven percent silver extraction had been obtained from the high-grade ore by a sulfurous acid treatment followed by cyanidation. However, this system would be costly as pretreatment with sulfur dioxide is an acid system, and extraction by cyanidation requires a basic system. To overcome this problem, a sulfurous acid-sodium chloride system was investigated. The chloride ion would act as a ligand and dissolve the silver as the tetrachloride silver complex (Ag Cl4 ³-). This reaction would be highly dependent on chloride ion concentration and would require 15 to 20 weight-percent sodium chloride to maintain solubility of 2,5 to 6.5 ounces of silver per ton of solution, as shown in figure 3. A solution containing 20-percent sodium chloride was used in all subsequent leaching experiments to insure solubility of the tetrachloro complex.
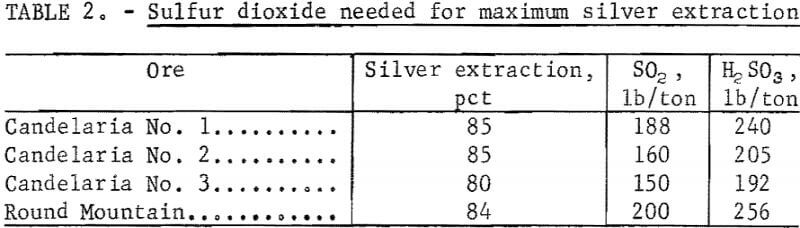
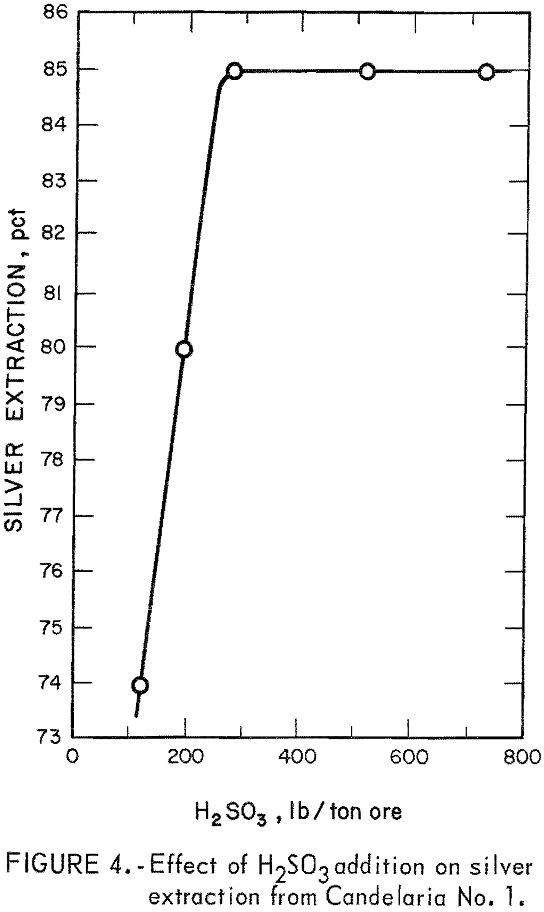
The first parameter investigated was the amount of SO2 required per ton of ore for maximum silver extraction. Sulfurous acid was used as a source of SO2 for these initial experiments.
The data in figure 4 show that silver extraction increased rapidly up to a maximum of 85 percent with increased additions of H2SO3 and then remained constant. The leveling off of silver extraction was attributed to the presence of argentojarosite mineral from which the sulfurous acid-sodium chloride leachant could not liberate and dissolve silver. Similar curves were obtained for other ores tested. The data in table 2 show that silver extractions of 80 to 85 percent were obtained from several ore samples using 150 to 240 pounds of SO2 per ton of ore.
Sulfurous acid for a large-scale plant operation would be obtained by burning sulfur in air to produce SO2. The SO2 would then be bubbled into ore slurry to form H2SO3. To determine whether there was any difference in silver extraction by bubbling SO2 into ore slurry as compared with adding H2SO3, a series of experiments was conducted in which the source of H2SO3 was SO2 bubbled into the pulp. Silver extractions were identical to the extractions obtained in earlier experiments. The amount of sulfur required for 80 to 85 percent silver extraction was 75 to 94 pounds per ton of ore.
The particle size of the ore is an important factor to be considered in any treatment sequence, because it not only influences the rate of reaction but it is directly related to liberating and exposing the mineral to the extractant. The effect of particle size on silver extraction from Candelaria No. 1 was investigated in a series of experiments grinding the ore from
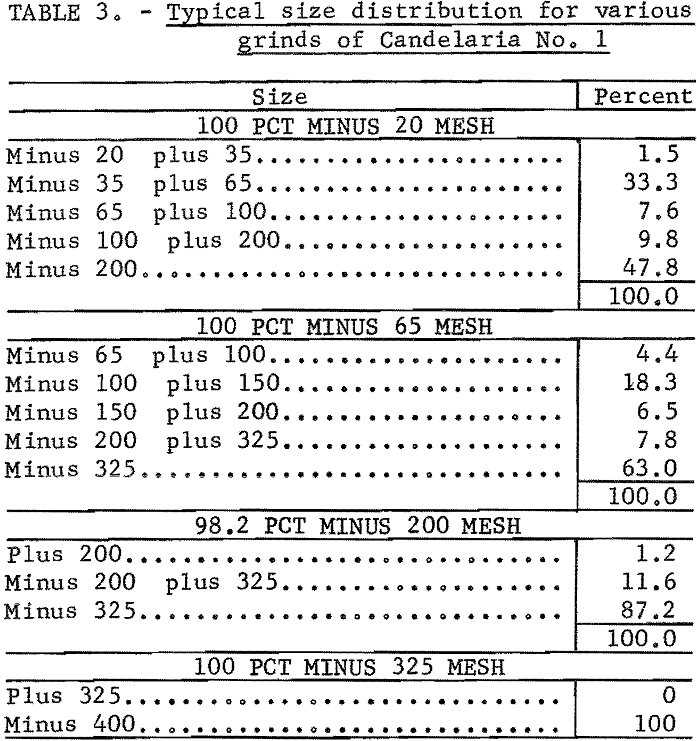
100 percent minus 20 mesh to 100 percent minus 400 mesh. Typical size distribution for several grinds is given in table 3.
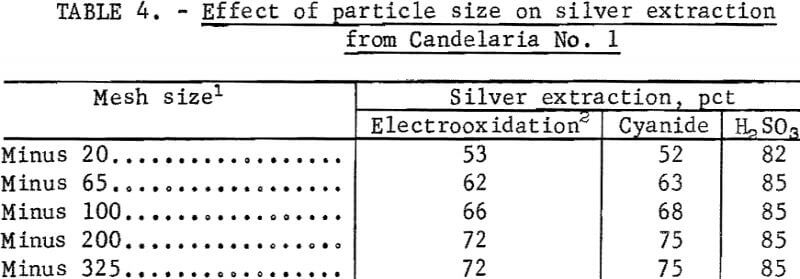
The effect of particle size on silver extraction from Candelaria No. 1 is shown in table 4. The data in table 4 indicate that silver extraction increased 19 percent and 23 percent, respectively, for electrooxidation and cyanidation after decreasing the particle size to 100 percent minus 200 mesh. Parallel experiments using the sulfurous acid-sodium chloride leach indicated that silver extraction was essentially independent of particle size for Candelaria No. 1 ore. This result was expected because the SO2-NaCl system dissolves the iron and manganese minerals containing the silver.
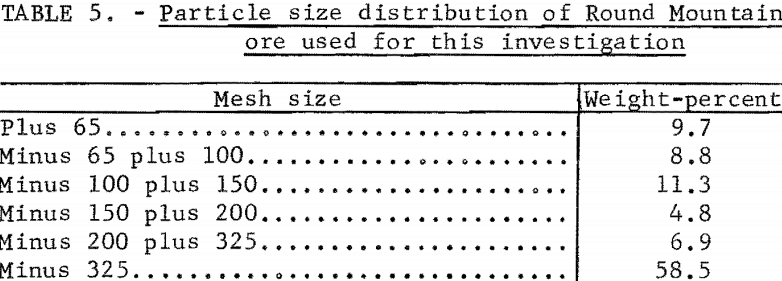
The Round Mountain ore reacted in a similar manner. The relatively coarse grind shown in table 5 was used for all experiments with the Round Mountain material.

An ore containing considerable amounts of silver encapsulated within silicious material, such as Candelaria No. 2 and No. 3, would not exhibit the same behavior as Candelaria No. 1 and Round Mountain ores. After grinding these more silicious ores to 100 percent minus 325 mesh, silver extraction by cyanidation treatment increased from 66 to 79 percent and 71 to 90 percent for Candelaria No. 2 and No. 3, respectively; corresponding silver extraction by sulfur dioxide-sodium chloride treatment increased from 85 to 93 percent and 80 to 90 percent, respectively, for the Candelaria No. 2 and No. 3 ores. However, fine grinding of ores is frequently impractical from an economic point of view.
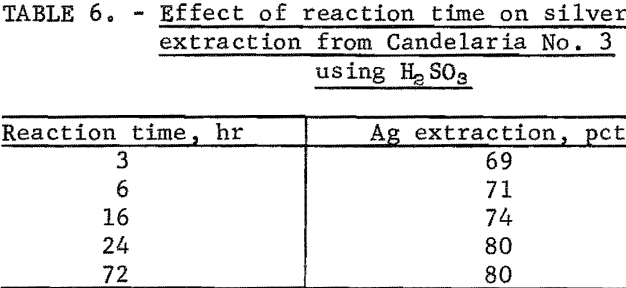
The rate of the dissolution of silver from the ores tested was very favorable, except for Candelaria No. 3. Optimum silver extraction was obtained within 2 hours from Candelaria No. 1, Candelaria No. 2, and Round Mountain in bottle tests using H2SO3. However, for Candelaria No. 3, maximum extraction was not reached until after a 24-hour reaction time. Table 6 shows the effect of reaction time on silver extraction from Candelaria No. 3. Silver extraction leveled off at 24 hours.

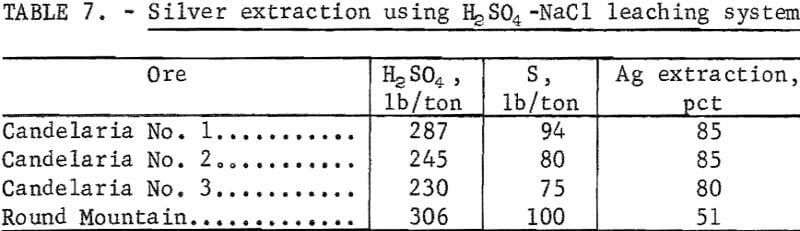
A comparative leach system substituting H2SO4 for the SO3 treatment was studied to determine the effect on silver extraction. The amount of H2SO4 used was based on the sulfur equivalent required for maximum extraction using the sulfur dioxide-sodium chloride system. The results of the H2SO4-NaCl leaching tests are shown in table 7. Silver extractions from the Candelaria ores were the same for sulfur dioxide-sodium chloride and sulfuric acid-sodium chloride. However, for the Round Mountain ore, silver extraction decreased from 84 to 51 percent with the use of H2SO4. Because much more of the silver in the Round Mountain ore occurs in manganese mineral than the Candelaria ores, sulfur dioxide-sodium chloride leach must be used for ores which contain substantial amounts of manganese.
Silver was recovered from the leach solutions generated in these experiments by precipitation using zinc or iron dust. The pH of the pregnant solution did not affect precipitation within a pH range of 2 to 7. Barren solutions containing less than 1 ppm silver were obtained. Silver metal was recovered from the precipitate by standard fire-refining techniques.
In comparing electrooxidation, cyanidation, and sulfur dioxide-sodium chloride leaching for limonitic ore, several facts are apparent from the data. For these ores, cyanidation and electrooxidation are comparable in terms of silver extraction. However, cyanide consumption for the ores tested was high, ranging from 3,8 to 9.0 pounds per ton. Higher silver extractions were consistently obtained using an acid-sodium chloride system. The chemistry involved in treating ores with sulfur dioxide-sodium chloride is very complex and large-scale testing is needed to evaluate the technique to determine its commercial applicability.
Conclusions
The refractory minerals in Candelaria and Round Mountain silver ore were identified, and their effects upon silver extraction were demonstrated. The data indicated that when silver is mineralogically associated with iron and manganese, a majority of these minerals must be dissolved or removed from the ore to obtain 80 to 85 percent silver extraction. Treatment with high concentrations of strong acid is required to liberate the remaining 15 to 20 percent silver from the argentojarosite mineral. Silver was extracted from these ores by applying a sulfurous acid-sodium chloride system. Silver extractions of 80 to 85 percent were obtained using 150 to 200 pounds of SO2 per ton of ore. For ores containing low amounts of silver in manganese(IV) minerals, sulfuric acid-sodium chloride leaching system can be substituted for the sulfur dioxide-sodium chloride system.
